Regulation of mitochondrial Turnover
Energy represents one of the most fundamental parameters in physiology and with respect to energy homeostasis, mitochondria are crucial organelle because they provide the majority of this energy under the form of adenosine triphosphate (ATP). To produce this ATP, mitochondria use different energy sources which are metabolized through a complex network. Together with this energy production, mitochondrial metabolism generates a variety of essential metabolites and it also controls homeostasis of signalling ions and molecules such as calcium, reactive oxygen species or neurotransmitters. Accordingly, mitochondria are pivotal structures for the cell metabolism but also for the global cell health.
Mitochondrial metabolic functions occur mainly in mitochondrial inner compartments: the mitochondrial matrix and inner membrane. These compartments are surrounded by a second membrane, the outer mitochondrial membrane. Despite this embedded localization, these metabolic activities must be constantly and finely orchestrated to fit the availability of substrates and the physiological needs.
Proteomic data from the literature as well as our own proteomic studies reveal that many mitochondrial metabolic proteins are ubiquitinylated. However, their implication in the regulation of the mitochondrial metabolism is still unknown. The aim of our project is to dissect the role played by the ubiquitin proteasome system UPS regarding the mitochondrial energy metabolism.
To achieve this aim, we have defined three specific objectives:
1. To characterize the mitochondrial ubiquitome and the UPS-dependent turnover of intra-mitochondrial proteins.
2. To identify and characterize mitochondrial E3 ubiquitin ligases.
3. To define the physiological consequences resulting from the UPS-dependent regulation of intramitochondrial proteins.
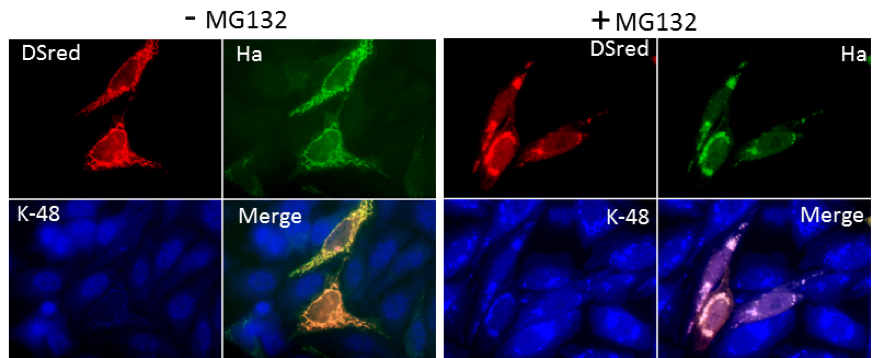
Accumulation of K48 ubiquitinated protein in mitochondria upon proteasome inhibition
Identification of mitochondrial E3s interactome by proteomics
Contributing Members
Giovanni Bénard [CNRS Director of Research, Principal Investigator]
Victor Gindensperger [Research Engineer]
Claude Lalou [Research Engineer]
Agata Ars [Technician] Adeola Folasade [Post Doc]
National and International Collaborations:
Patryk Ngondo [Institut de Biologie Moléculaire des Plantes, Strasbourg, France]
Rajesh Singh [University of Baroda, India]
Timothy Shutt [University of Calgary, Canada]
Jean-William Dupuy [Proteomics Facility, University of Bordeaux, France]
METATOUL [INSA, INRAe, Toulouse]
Selected publications:
Lavie J, Lalou C, Mahfouf W, Dupuy JW, Lacaule A, Cywinska AA, Lacombe D, Duchêne AM, Raymond AA, Rezvani HR, Ngondo RP, Bénard G. The E3 ubiquitin ligase FBXL6 controls the quality of newly synthesized mitochondrial ribosomal proteins. Cell Rep. 2023 Jun 27;42(6):112579. Epub 2023 Jun 1. DOI: 10.1016/j.celrep.2023.112579
Lavie J, De Belvalet H, Sonon S, Ion AM, Dumon E, Melser S, Lacombe D, Dupuy JW, Lalou C, Bénard G. Ubiquitin-Dependent Degradation of Mitochondrial Proteins Regulates Energy Metabolism. Cell Report. 2018.
Hebert-Chatelain E, Desprez T, Serrat R, Bellocchio L, Soria-Gomez E, Busquets-Garcia A, Pagano Zottola AC, Delamarre A, Cannich A, Vincent P, Varilh M, Robin LM, Terral G, García-Fernández MD, Colavita M, Mazier W, Drago F, Puente N, Reguero L, Elezgarai I, Dupuy JW, Cota D, Lopez-Rodriguez ML, Barreda-Gómez G, Massa F, Grandes P, Bénard G*, Marsicano G*. A cannabinoid link between mitochondria and memory. Nature. 2016. [*Sharing senior authorship]
Lavie J, Serrat R, Bellance N, Courtand G, Dupuy JW, Tesson C, Coupry I, Brice A, Lacombe D, Durr A, Stevanin G, Darios F, Rossignol R, Goizet C, Bénard G. Mitochondrial morphology and cellular distribution are altered in SPG31 patients and are linked to DRP1 hyperphosphorylation. Hum. Mol. Genet. 2017.
Tomar D, Prajapati P, Lavie J, Singh K, Lakshmi S, Bhatelia K, Roy M, Singh R, Singh R*, Bénard G*. TRIM4; a novel mitochondrial interacting RING E3 ligase, sensitizes the cells to hydrogen peroxide (H2O2) induced cell death. Free Radic Biol Med. 2015. [*Sharing senior authorship]
Melser S, Hebert Chatelain E, Lavie J, Mahfouf W, Jose C, Obre E, Goorden S, Priaul M, Elgersma Y, Rezvan H, Rossignol R, Bénard G. Rheb regulates mitophagy induced by mitochondrial energetic status. Cell Metabolism. 2013.
